Summary Of Research Goals
The research in the Johnson laboratory focuses on the roles of heparan sulfate proteoglycans (HSPGs) and chondroitin sulfate proteoglycans (CSPGs) in central nervous system development in Drosophila. Proteoglycans are evolutionary ancient molecules whose roles in development are only now beginning to be addressed. At present, we are focusing our efforts on one specific HSPG, called Syndecan, which we have found to have significant defects in the developing nervous system involving both axon guidance and synapse formation. We are currently at an extremely rare and fortunate posltion on these projects- we have robust and highly penetrant phenotypes in several aspects of nervous system development, and we have all the reagents and equipment to make significant progress on these research proposals. We are also carrying out biochemical and genetic approaches to identify CSPGs in Drosophila.
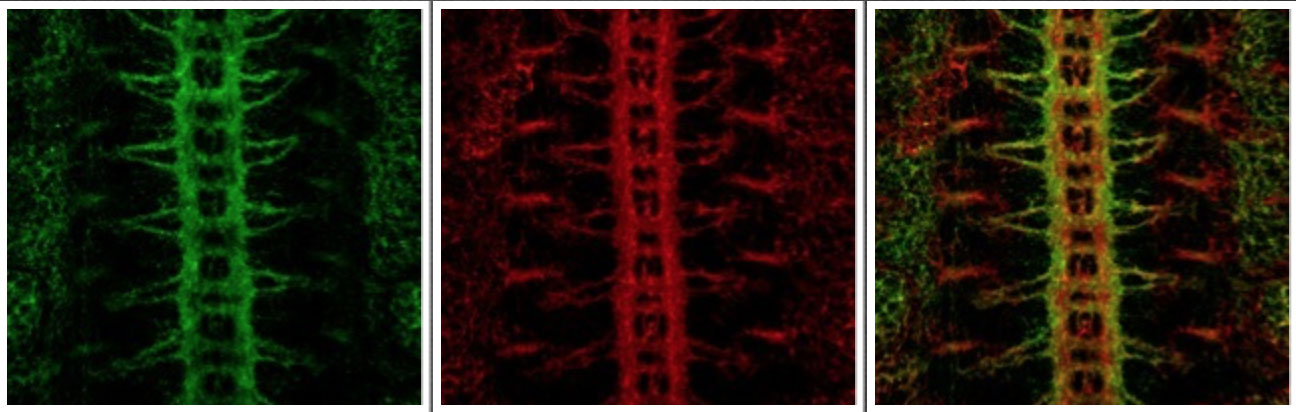
1. HSPGs and guidance at the central nervous system midline
In the developing Drosophila embryo (as in all bilaterally symmetric organisms) the vast majority of neurons on one side of the CNS send projections that cross the midline once to reach targets on the opposite side of the body. To do this, neurons must send out processes that cross the midline of the CNS once, and only once. Several genes have been identified as lkey players in controlling neuronal process outgrowth at the midline of the CNS, including the repulsive molecule Slit and its receptor Robo. After a neuronal process crosses the midline, it begins expressing the Robo receptor, becomes sensitive to the repellant Slit, and the process will not cross the midline again.
In Syndecan mutants (Sdc), we found a surprising mutant phenotype in which neuronal processes cross and recross the midline, apparently insensitive to the repellant Slit (despite expressing Robo receptors). In Syndecan mutant embryos, approximately 40% of segments have defects in midline crossing. Further studies demonstrated much to our surprise, that in the absence of heparan sulfate, Slit and Robo do not bind to one another. Rather, heparan sulfate forms an essential molecular bridge between Slit and Robo, and is a crucial player in controlling midline guidance through its ability to link Slit and Robo into a function complex (Johnson et al., 2003). Further studies in my lab have shown that mutations in another HSPG, called Dallylike, have no midline guidance defects, but instead have defects in guidance away from the midline. Double mutant embryos lacking both Syndecan and Dallylike, however, have much more severe phenotypes than either mutant alone. In these double mutant embryos, all neuronal processes are attracted to the midline and are nearly insensitive to the repellant Slit (Smart et al., 2011).
2. HSPGs and the control of synapse morphogenesis
Sdc has been shown to control synapse formation in Drosophila (Johnson et al, 2006). While we know that Sdc acts with the receptor protein LAR to increase the size of the synapse, we do not know the mechanism by which these proteins promote synapse growth. Sdc could function simply as a ligand for LAR, or Sdc could serve as an instructive receptor that is itself responsible for initiating the growth of the synapse. The second major project in my lab will attempt to identify which of these models is correct. Sdc is a transmembrane protein, and the cytoplasmic domains of Sdc are highly conserved across species. Comparisons between Drosophila Sdc and the human homologues of Sdc reveal 60-80% sequence identity in the cytoplasmic domains. This high degree of sequence identity suggests a conserved role for Sdc between Drosophila and humans. The function of these domains, however, has remained elusive in all organisms. Recently, we generated a variety of Sdc mutant expression constructs to identify which parts of Sdc are required for its function. Using expression constructs that lack the cytoplasmic domains, the extracellular domains, and the heparan sulfate attachment sites, we demonstrated that both the cytoplasmic and extracellular domains of Syndecan are required for Sdc to promote synapse growth at the larval NMJ (Nguyen et al., 2016)
3. The identification of CSPGs in Drosophila
Chondroitin sulfate proteoglycans (CSPGs) are a diverse family of extracellular matrix proteins that consist of a core protein with covalently attached chondroitin sulfate (CS) sidechains. CSPGs show high levels of expression, and play crucial roles as repulsive guidance cues, during many aspects of central nervous system development. However, CSPG function is not limited to the developing CNS – roles for CSPGs are also well-established on glial scars at sites of CNS injury, and in the regulation of synaptic plasticity during critical period closure. The vast majority of research on CSPGs has been conducted in vertebrate systems. In fact, although CSPGs have recently been identified in C. elegans, there are no bona fide CSPGs in Drosophila. This lack of identified CSPGs in Drosophila is particularly surprising since CSPGs are found in both nematodes and chordates, and that the arthropod lineage diverged from chordates later than nematodes. The lack of identified CSPGs in Drosophila suggests that either CSPGs have been lost from the Drosophila genome, or that Drosophila have CSPGs that have yet to be identified.
There are three lines of evidence suggesting that Drosophila have CSPGs. First, the enzymes involved in CS synthesis and polymerization are highly conserved between species, including Drosophila. Second, the disaccharides that make up chondroitin sulfate (N-acetyl-galactosamine and glucuronic acid) are found in high concentrations in embryonic, larval and adult Drosophila tissues. Third, when an HSPG like Sdc is overexpressed in Drosophila embryos, it carries both HS and CS sidechains – even though the endogenously expressed Sdc is exclusively an HSPG. Together these data provide strong circumstantial evidence that CSPGs exist, but haven’t yet been identified in Drosophila.
We propose to identify the first Drosophila CSPGs, and to examine the roles of CSPGs and CSPG receptors in Drosophila nervous system development by exploring the following questions:
a. What genes in Drosophila represent CSPGs?
Unlike HSPG core proteins that are conserved between species, outside of the CS attachment site of SGxG, CSPG core proteins tend to have very weak sequence similarity across species and are not readily identifiable with bioinformatics approaches. To identify CSPGs, we will pursue a biochemical approach that was used to successfully identify the first chondroitin proteoglycans (CPGs) in C. elegans: Large-scale isolation of larval protein extracts, followed by anion exchange chromatography and mass spectrometry.
b. What roles do CS and CSPGs play during Drosophila development?
CS biosynthetic enzymes are highly conserved between species. We have identified the Drosophila homologues of several CS biosynthetic enzymes, none of which have known mutant phenotypes. RNAi lines exist for these genes, allowing us to knock down CS biosynthesis in a spatially and temporally controlled manner. In addition to RNAi-mediated knockdown, we can generate loss of function alleles of these genes using either P-element or CRISPr-mediated mutagenesis. To examine the roles of individual CSPGs during development, similar mutagenesis strategies will be employed to knock out the CSPGs identified. Expression constructs will be made for identified CSPGs and, using the Gal4-UAS system, we will express CSPG core proteins to probe their roles in development.
c. What are the receptors for CS and CSPGs in Drosophila?
In order to generate a more complete picture for how CS and CSPGs regulate nervous system development, we will screen the Drosophila extracellular interactome for proteins that bind to HS and CS. Using biotinylated CS and HS attached to the surface of 96 well plates, we will screen the 202 AP-fusion proteins in the interactome for differential binding to HS and CS. This interactome has an internal positive control; Drosophila LAR is a member of the interactome and is known to bind to HS with nanomolar affinity and the vertebrate homologues bind both HS and CS. We predict that Drosophila LAR will bind CS and anticipate finding several additional CS-binding proteins using this assay.
Funding
Research in the Johnson lab has been funded by the following grants:
Hirsch Research Initiation Grant 4/2013-6/2015 $10,000 for the Identification of Novel Chondroitin Sulfate Proteoglycans in Drosophila
National Science Foundation- Research at Undergraduate Institutions 9/2009 – 8/2013
Division of Integrative Organismal Systems – Neural Systems Cluster
“Investigating the mechanisms of Syndecan function during nervous system development”
Grant# 0841551
National Institutes of Health – Academic Research Enhancement Award 5/2009- 8/2009
National Institute of Neurological Disorders and Stroke
“Characterizing the function of Syndecan during CNS development”
Grant # 1 R15NS065433-01